 Their sophisticated detection algorithm easily and precisely identifies plasmid features that differ by only a few nucleotides, such as the fluorescent proteins EGFP and mEGFP. SnapGene’s Feature Library - SnapGene’s software seamlessly identifies a variety of common plasmid features such as antibiotic resistance genes, selectable markers, promoters,origins,tags, and certain ORFs. These practical and effective changes will give you the ability to find the plasmid with the features you need quickly. Three factors make our SnapGene powered plasmid maps far easier to navigate. 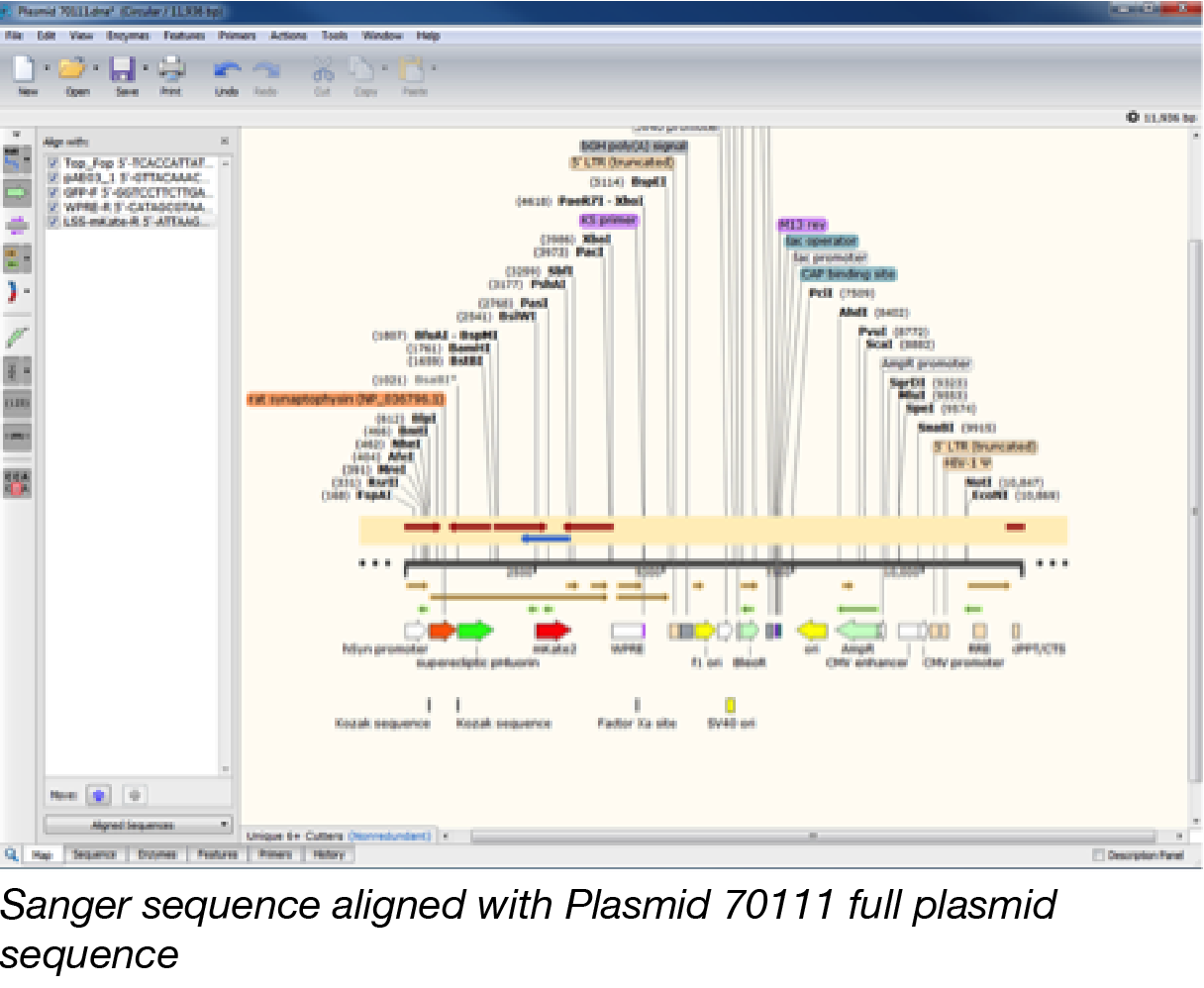
Overview of SnapGene Powered Plasmid Maps at Addgene We'll be monitoring and improving the maps further after this initial launch so stay tuned for more updates. Read on to learn more about the improvements we’ve implemented and how they’ll make it easier for you to find the plasmids you need. Our old map is on the left while the SnapGene powered map is on the right ( click here to see the new map in action). 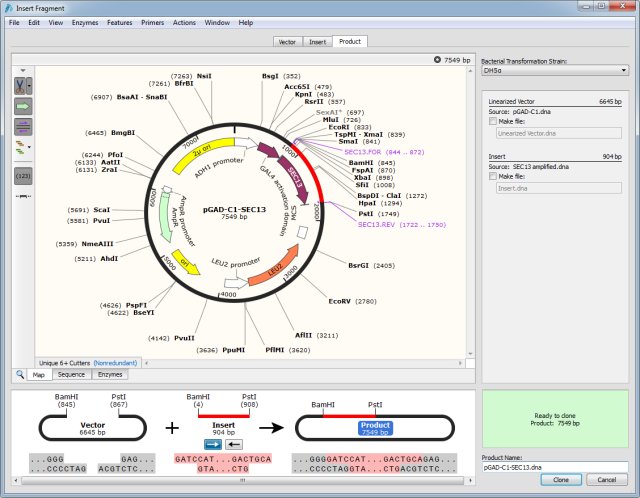
For a quick look at just how much things have improved, check out the example below. With the backing of SnapGene’s sequence viewer software and extensive feature library, our updated plasmid and sequence displays are now much easier to interpret and analyze at a glance. *.zip, *.7z, *.gzip, *.gz, *.bz2, *.tar, *.In meetings, in surveys, on Twitter - there is one thing we've heard over and over from our users: " Please, please improve your plasmid maps!" After thoughtful design, vetting, and tweaking, we’re excited to announce that our plasmid and sequence displays are now powered by GSL Biotech's SnapGene Server Software. Table 1: File Types Imported by SeqMan NGen for De Novo Assemblies and Templated Alignments (Special Workflows) Note: SeqMan NGen supports the export of data in. Below are the file types supported for import for both normal workflows and special workflows. 
SeqMan NGen is included within Lasergene Genomics. Image Files (*.png, *.jpg, *.gif, *.bmp 8)ġ File-Save will save back as original file type. These files are not supported through the menu system.ĩ Projects or files must contain protein sequence data.ġ0 Only the sequence, features and comments of the file will be read in.ġ3 Lasergene does not support ABI files without base calls.ġ4 Protean 3D supports version 3.2 of PDB files.ġ5 Available to support gap closure in SeqMan Ultra and for assembly in SeqMan NGen.ġ6 Supports *.txt and *.gp UniProt files only.įASTA Format (*.fas, *.fap, *.fna, *.fasta) All other files will be ignored.Ģ Multiple Lasergene DNA sequence files can be created by EditSeq, SeqMan Pro, and MegAlign.ģ FOF can not contain reference to another multiple sequence file format such as FastA, DNA* mseq, or Zip.ĥ Referred to collectively as “Lasergene Documents” in “Files of Type” or “Show pull-down menus”.ħ GenBank formatted features will be parsed.Ĩ Through “drag and drop” only. SeqMan NGen Transcriptome Files (*.transcriptome)ġ Only zip files containing ABI, SCF3, Lasergene protein or Lasergene DNA files. SeqMan NGen Assembly Projects (*.assembly) Protein Data Bank files (*.pdb, *.ent, *.pdb.gz, *.ent.gz, *.zip, *.txt) 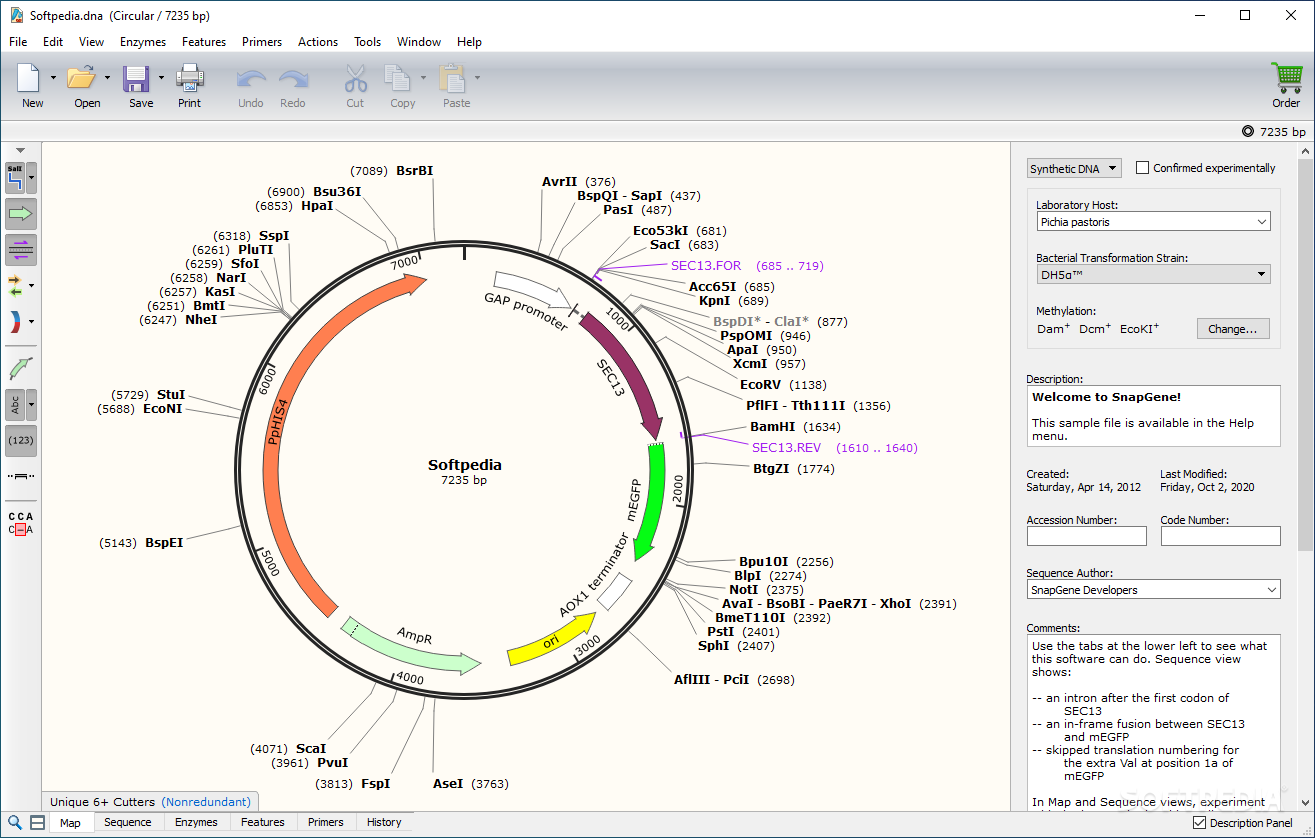
Imported File Types File FormatĤ54 Life Sciences output files (*.sff *.fas and *.qual)įASTA Format (*.fasta, *.fas, *.fap, *.nt, *.aa)įASTA Alignment with Gaps (*.fasta, *.fa, *.fas, *.fna, *.fap, *.nt, *.aa, *.frn, *.faa, *.ffn, *.mfa) Lasergene Molecular Biology and Lasergene Proteinīelow are the file formats supported for import and export by the applications within the Lasergene Molecular Biology and Lasergene Protein packages. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
